Development of a parameter-efficient method for biomedical image synthesis by substituting text conditioning with pathology foundation model embeddings in latent diffusion
DOI:
https://doi.org/10.15587/2706-5448.2026.355663Keywords:
latent diffusion models, pathology foundation models, histopathology image synthesis, medical image generationAbstract
The object of research is the process of synthesizing patches of histopathological images conditioned by embeddings of the pathology foundation model. One of the key problems is that existing approaches to diffusion synthesis either rely on text conditioning via CLIP encoders, which lack morphological understanding, or require full retraining of the generative base model, which requires significant computational resources.
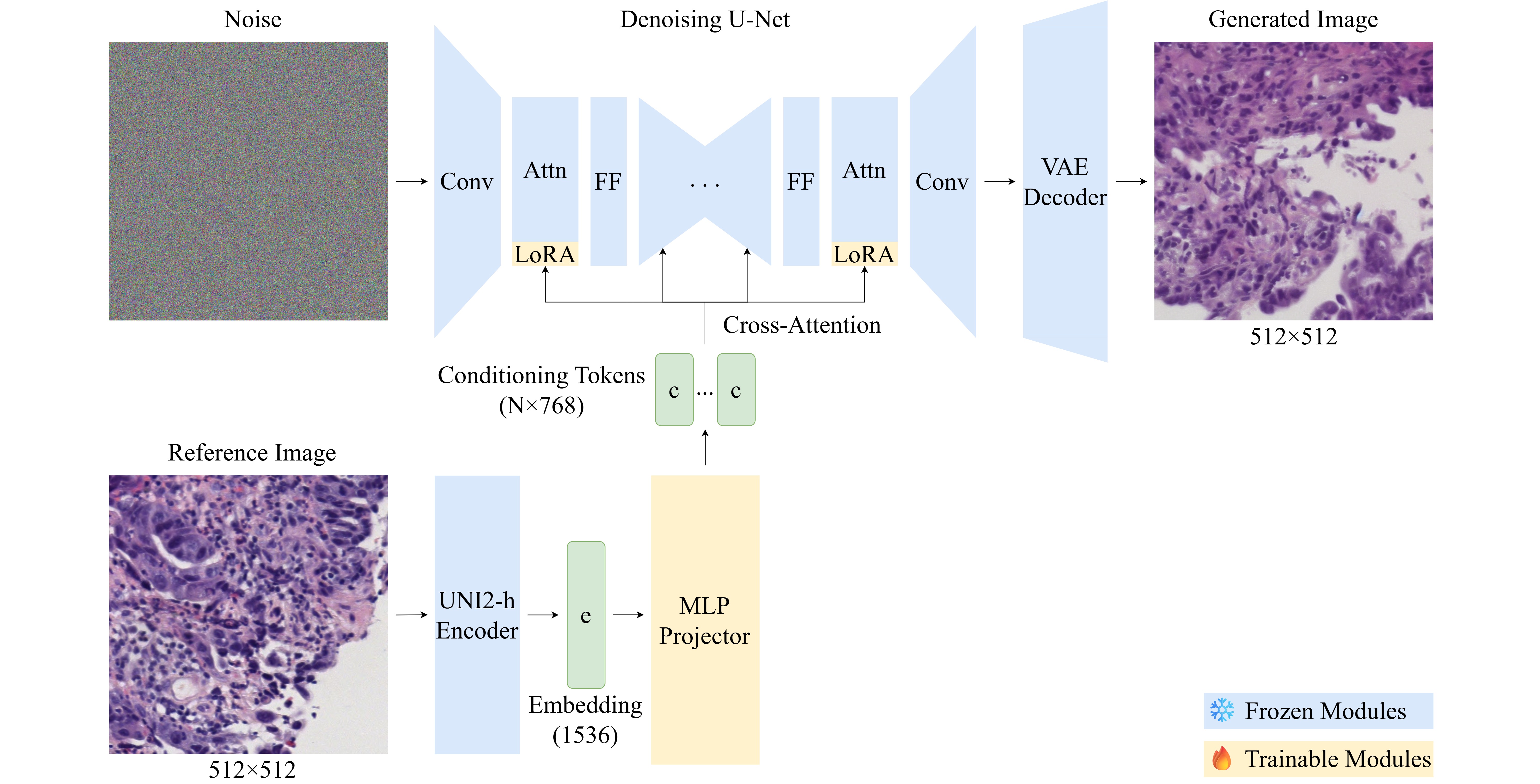
The research used a parameter-efficient adaptation of the previously trained latent diffusion model using low-rank adaptation (LoRA) of the U-Net attention layers in combination with a training MLP projector that reflects the embeddings of the pathology foundation model UNI2-h in the conditioning space of the cross-attention mechanism. Ablation studies of 12 configurations were conducted varying the adapter rank, the number of conditioning tokens, and the projector architecture.
It is confirmed that embeddings of the pathology foundation model can effectively replace text conditioning for the synthesis of histopathology images in a parameter-efficient mode. The optimal configuration achieved FID 77.59 on the validation set and FID 84.17 on the test set when training only 5.53 million parameters, which is 0.64% of the parameters of the base model. This is due to the fact that the proposed method has a number of characteristic features, in particular: embeddings of the pathology foundation model provide morphologically richer conditioning than CLIP-based text representations, and low rank adaptation limits the trainable space to the conditioning pathway.
This provides the possibility of generating histopathology images without text annotations and without full retraining of the model using approximately 12 GB of video memory. Compared to the previous text-conditioned approach on the same dataset, which demonstrated class-wise FID values in the range of 113 to 138, the embedding conditioning method provides significantly higher generation quality while maintaining parameter efficiency.
References
- Litjens, G., Bandi, P., Ehteshami Bejnordi, B., Geessink, O., Balkenhol, M., Bult, P. et al. (2018). 1399 H&E-stained sentinel lymph node sections of breast cancer patients: the CAMELYON dataset. GigaScience, 7 (6). https://doi.org/10.1093/gigascience/giy065
- Walsh, E., Orsi, N. M. (2024). The current troubled state of the global pathology workforce: a concise review. Diagnostic Pathology, 19 (1). https://doi.org/10.1186/s13000-024-01590-2
- Guan, H., Yap, P.-T., Bozoki, A., Liu, M. (2024). Federated learning for medical image analysis: A survey. Pattern Recognition, 151, 110424. https://doi.org/10.1016/j.patcog.2024.110424
- Zhang, Y., Kang, B., Hooi, B., Yan, S., Feng, J. (2023). Deep Long-Tailed Learning: A Survey. IEEE Transactions on Pattern Analysis and Machine Intelligence, 45 (9), 10795–10816. https://doi.org/10.1109/tpami.2023.3268118
- Campanella, G., Hanna, M. G., Geneslaw, L., Miraflor, A., Werneck Krauss Silva, V., Busam, K. J. et al. (2019). Clinical-grade computational pathology using weakly supervised deep learning on whole slide images. Nature Medicine, 25 (8), 1301–1309. https://doi.org/10.1038/s41591-019-0508-1
- Jose, L., Liu, S., Russo, C., Nadort, A., Di Ieva, A. (2021). Generative Adversarial Networks in Digital Pathology and Histopathological Image Processing: A Review. Journal of Pathology Informatics, 12 (1), 43. https://doi.org/10.4103/jpi.jpi_103_20
- Saad, M. M., O’Reilly, R., Rehmani, M. H. (2024). A survey on training challenges in generative adversarial networks for biomedical image analysis. Artificial Intelligence Review, 57 (2). https://doi.org/10.1007/s10462-023-10624-y
- Dhariwal, P., Nichol, A. (2021). Diffusion models beat GANs on image synthesis. arXiv:2105.05233. https://doi.org/10.48550/arXiv.2105.05233
- Rombach, R., Blattmann, A., Lorenz, D., Esser, P., Ommer, B. (2022). High-Resolution Image Synthesis with Latent Diffusion Models. 2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 10674–10685. https://doi.org/10.1109/cvpr52688.2022.01042
- Radford, A., Kim, J. W., Hallacy, C., Ramesh, A., Goh, G., Agarwal, S. et al. (2021). Learning transferable visual models from natural language supervision. arXiv:2103.00020. https://doi.org/10.48550/arXiv.2103.00020
- Chen, R. J., Ding, T., Lu, M. Y., Williamson, D. F. K., Jaume, G., Song, A. H. et al. (2024). Towards a general-purpose foundation model for computational pathology. Nature Medicine, 30, 850–862. https://doi.org/10.1038/s41591-024-02857-3
- Yellapragada, S., Graikos, A., Prasanna, P., Kurc, T., Saltz, J., Samaras, D. (2024). PathLDM: Text conditioned Latent Diffusion Model for Histopathology. 2024 IEEE/CVF Winter Conference on Applications of Computer Vision (WACV), 5170–5179. https://doi.org/10.1109/wacv57701.2024.00510
- Graikos, A., Yellapragada, S., Le, M.-Q., Kapse, S., Prasanna, P., Saltz, J., Samaras, D. (2024). Learned Representation-Guided Diffusion Models for Large-Image Generation. 2024 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 8532–8542. https://doi.org/10.1109/cvpr52733.2024.00815
- Boada, J. C., Umer, R. M., Marr, C. (2025). CytoDiff: AI-Driven Cytomorphology Image Synthesis for Medical Diagnostics. 2025 IEEE/CVF International Conference on Computer Vision Workshops (ICCVW), 1136–1144. https://doi.org/10.1109/iccvw69036.2025.00122
- Hu, E. J., Shen, Y., Wallis, P., Allen-Zhu, Z., Li, Y., Wang, S. et al. (2022). LoRA: low-rank adaptation of large language models. arXiv:2106.09685. https://doi.org/10.48550/arXiv.2106.09685
- Ho, J., Jain, A., Abbeel, P. (2020). Denoising diffusion probabilistic models. Proceedings of the 34th International Conference on Neural Information Processing Systems, 34, 6840–6851. https://doi.org/10.48550/arXiv.2006.11239
- Yellapragada, S., Graikos, A., Triaridis, K., Prasanna, P., Gupta, R., Saltz, J., Samaras, D. (2025). ZoomLDM: Latent Diffusion Model for multi-scale image generation. 2025 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 23453–23463. https://doi.org/10.1109/cvpr52734.2025.02184
- Mao, Y., Li, H., Pang, W., Papanastasiou, G., Yang, G., Wang, C. (2024). SeLoRA: self-expanding low-rank adaptation of latent diffusion model for medical image synthesis. arXiv:2408.07196. https://doi.org/10.48550/arXiv.2408.07196
- Berezsky, O., Melnyk, G., Liashchynskyi, P., Pitsun, O.; Babichev, S., Lytvynenko, V. (Eds.) (2025). Biomedical Image Datasets. Lecture Notes on Data Engineering and Communications Technologies, vol 244. Cham: Springer, 61–82. https://doi.org/10.1007/978-3-031-88483-2_3
- Berezsky, O., Liashchynskyi, P., Melnyk, G., Dombrovskyi, M., Berezkyi, M. (2024). Synthesis of biomedical images based on generative intelligence tools. Proceedings of the 7th International Conference on Informatics & Data-Driven Medicine (IDDM 2024). Birmingham. CEUR Workshop Proceedings, 3892, 349–362. Available at: https://ceur-ws.org/Vol-3892/paper23.pdf
- Berezsky, O., Liashchynskyi, P., Pitsun, O., Izonin, I. (2024). Synthesis of Convolutional Neural Network architectures for biomedical image classification. Biomedical Signal Processing and Control, 95, 106325. https://doi.org/10.1016/j.bspc.2024.106325
- Berezsky, O., Liashchynskyi, P., Pitsun, O., Melnyk, G. (2024). Method and Software Tool for Generating Artificial Databases of Biomedical Images Based on Deep Neural Networks. 6th International Conference on Informatics & Data-Driven Medicine Bratislava. https://doi.org/10.48550/arXiv.2405.16119
- Kuzmin, S., Berezsky, O. (2025). Analysis of diffusion models and biomedical image generation tools. Computer Systems and Information Technologies, 2, 8–19. https://doi.org/10.31891/csit-2025-2-1
- Zhu, C., Chen, W., Peng, T., Wang, Y., Jin, M. (2022). Hard Sample Aware Noise Robust Learning for Histopathology Image Classification. IEEE Transactions on Medical Imaging, 41 (4), 881–894. https://doi.org/10.1109/tmi.2021.3125459
- Ho, J., Salimans, T. (2022). Classifier-free diffusion guidance. arXiv:2207.12598. https://doi.org/10.48550/arXiv.2207.12598
- Heusel, M., Ramsauer, H., Unterthiner, T., Nessler, B., Hochreiter, S. (2017). GANs trained by a two time-scale update rule converge to a local Nash equilibrium. 31st Conference on Neural Information Processing Systems (NIPS 2017). Long Beac. https://doi.org/10.48550/arXiv.1706.08500
- Bińkowski, M., Sutherland, D. J., Arbel, M., Gretton, A. (2018). Demystifying MMD GANs. International Conference on Learning Representations (ICLR). https://doi.org/10.48550/arXiv.1801.01401
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2026 Sergii Kuzmin, Oleh Berezsky

This work is licensed under a Creative Commons Attribution 4.0 International License.
The consolidation and conditions for the transfer of copyright (identification of authorship) is carried out in the License Agreement. In particular, the authors reserve the right to the authorship of their manuscript and transfer the first publication of this work to the journal under the terms of the Creative Commons CC BY license. At the same time, they have the right to conclude on their own additional agreements concerning the non-exclusive distribution of the work in the form in which it was published by this journal, but provided that the link to the first publication of the article in this journal is preserved.







