Improvement of the method of the multiclass Pap smear image segmentation based on cross-domain transfer learning with limited data
DOI:
https://doi.org/10.15587/1729-4061.2026.352892Keywords:
transfer learning, Pap smear, cervical cancer, segmentation, deep learningAbstract
This study examined automated multi-class semantic segmentation of Pap smear images used for cervical cancer detection. The effectiveness of existing deep learning methods is often limited due to a lack of labeled data, high morphological variability of cervical cells, overlapping structures, noise, low contrast, and imaging artifacts characteristic of cytology specimens.
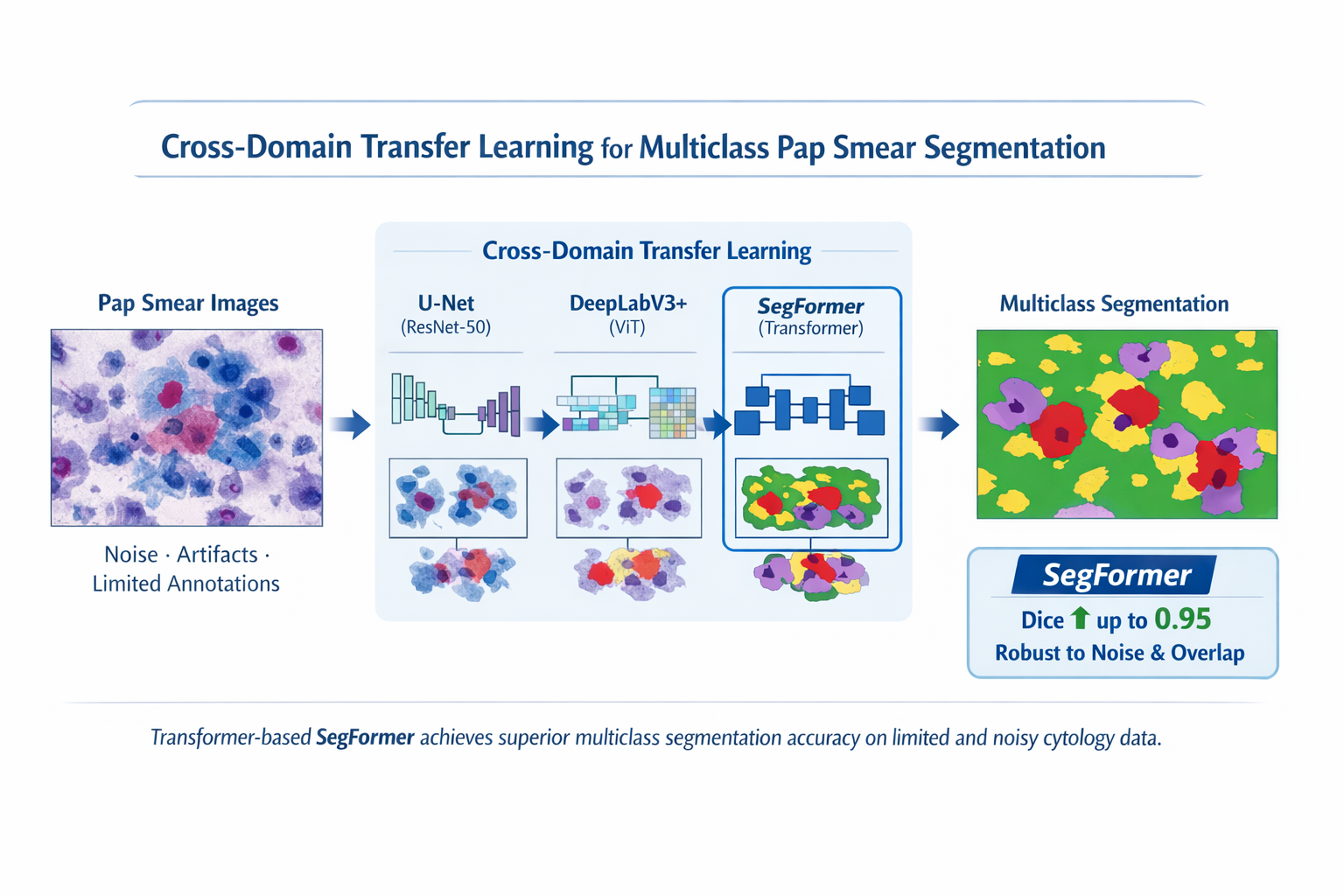
In this study, the authors propose a cross-domain transfer learning approach that adapts pre-trained deep neural networks to the task of multi-class Pap smear segmentation. All networks were pre-trained on large-scale natural image datasets. In the experiments, both convolutional neural networks and Transformer-based models, including hybrid configurations, were refined and systematically compared. Network performance was assessed using quantitative metrics (Dice score, IoU, HD95), as well as qualitative visual assessment of segmentation edges and boundaries.
The results obtained from the experiments showed that Transformer-based architectures, in particular SegFormer, significantly outperform convolutional models when processing noisy and heterogeneous cytological data. Using specialized data augmentation strategies developed specifically for medical imaging, SegFormer increased Dice scores to 0.95 across all classes (healthy, unhealthy, rubbish, both cells), as well as improved edge accuracy and robustness to artifacts and cell aliasing.
Multi-scale feature extraction and global context modeling proved essential for accurately identifying cellular structures in data-constrained settings. The results obtained in the study can help in the development of reliable automated diagnostic tools to assist cytopathologists, as well as to improve the overall accuracy and efficiency of cervical cancer screening programs
References
- Mustafa, W. A., Ismail, S., Mokhtar, F. S., Alquran, H., Al-Issa, Y. (2023). Cervical Cancer Detection Techniques: A Chronological Review. Diagnostics, 13 (10), 1763. https://doi.org/10.3390/diagnostics13101763
- Litjens, G., Kooi, T., Bejnordi, B. E., Setio, A. A. A., Ciompi, F., Ghafoorian, M. et al. (2017). A survey on deep learning in medical image analysis. Medical Image Analysis, 42, 60–88. https://doi.org/10.1016/j.media.2017.07.005
- Fang, M., Liao, B., Lei, X., Wu, F.-X. (2024). A systematic review on deep learning based methods for cervical cell image analysis. Neurocomputing, 610, 128630. https://doi.org/10.1016/j.neucom.2024.128630
- Cruz-Roa, A., Basavanhally, A., González, F., Gilmore, H., Feldman, M., Ganesan, S. et al. (2014). Automatic detection of invasive ductal carcinoma in whole slide images with convolutional neural networks. Medical Imaging 2014: Digital Pathology, 9041, 904103. https://doi.org/10.1117/12.2043872
- Pati, P., Jaume, G., Foncubierta-Rodríguez, A., Feroce, F., Anniciello, A. M., Scognamiglio, G. et al. (2022). Hierarchical graph representations in digital pathology. Medical Image Analysis, 75, 102264. https://doi.org/10.1016/j.media.2021.102264
- Win, K. Y., Choomchuay, S. (2017). Automated segmentation of cell nuclei in cytology pleural fluid images using OTSU thresholding. 2017 International Conference on Digital Arts, Media and Technology (ICDAMT), 14–18. https://doi.org/10.1109/icdamt.2017.7904925
- Ronneberger, O., Fischer, P., Brox, T. (2015). U-Net: Convolutional Networks for Biomedical Image Segmentation. Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015, 234–241. https://doi.org/10.1007/978-3-319-24574-4_28
- Oktay, O., Schlemper, J., Folgoc, L. L., Lee, M., Heinrich, M., Misawa, K. et al. (2018). Attention U-Net: Learning Where to Look for the Pancreas. arXiv. https://doi.org/10.48550/arXiv.1804.03999
- Hatamizadeh, A., Tang, Y., Nath, V., Yang, D., Myronenko, A., Landman, B. et al. (2022). UNETR: Transformers for 3D Medical Image Segmentation. 2022 IEEE/CVF Winter Conference on Applications of Computer Vision (WACV), 1748–1758. https://doi.org/10.1109/wacv51458.2022.00181
- Chen, J., Lu, Y., Yu, Q., Luo, X., Adeli, E., Wang, Y. et al. (2021). TransUNet: Transformers Make Strong Encoders for Medical Image Segmentation. arXiv. https://doi.org/10.48550/arXiv.2102.04306
- Wang, W., Xie, E., Li, X., Fan, D.-P., Song, K., Liang, D. et al. (2021). Pyramid Vision Transformer: A Versatile Backbone for Dense Prediction without Convolutions. 2021 IEEE/CVF International Conference on Computer Vision (ICCV), 548–558. https://doi.org/10.1109/iccv48922.2021.00061
- Xu, C., Li, M., Li, G., Zhang, Y., Sun, C., Bai, N. (2022). Cervical Cell/Clumps Detection in Cytology Images Using Transfer Learning. Diagnostics, 12 (10), 2477. https://doi.org/10.3390/diagnostics12102477
- Raghu, M., Zhang, C., Kleinberg, J., Bengio, S. (2019). Transfusion: Understanding Transfer Learning for Medical Imaging. arXiv. https://doi.org/10.48550/arXiv.1902.07208
- Tajbakhsh, N., Shin, J. Y., Gurudu, S. R., Hurst, R. T., Kendall, C. B., Gotway, M. B., Liang, J. (2016). Convolutional Neural Networks for Medical Image Analysis: Full Training or Fine Tuning? IEEE Transactions on Medical Imaging, 35 (5), 1299–1312. https://doi.org/10.1109/tmi.2016.2535302
- He, K., Zhang, X., Ren, S., Sun, J. (2016). Deep Residual Learning for Image Recognition. 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 770–778. https://doi.org/10.1109/cvpr.2016.90
- Russakovsky, O., Deng, J., Su, H., Krause, J., Satheesh, S., Ma, S. et al. (2015). ImageNet Large Scale Visual Recognition Challenge. International Journal of Computer Vision, 115 (3), 211–252. https://doi.org/10.1007/s11263-015-0816-y
- Dosovitskiy, A., Beyer, L., Kolesnikov, A., Weissenborn, D., Zhai, X., Unterthiner, T. et al. (2020). An Image is Worth 16x16 Words: Transformers for Image Recognition at Scale. arXiv. https://doi.org/10.48550/arXiv.2010.11929
- Chen, L.-C., Zhu, Y., Papandreou, G., Schroff, F., Adam, H. (2018). Encoder-Decoder with Atrous Separable Convolution for Semantic Image Segmentation. Computer Vision – ECCV 2018, 833–851. https://doi.org/10.1007/978-3-030-01234-2_49
- Xie, E., Wang, W., Yu, Z., Anandkumar, A., Alvarez, J. M., Luo, P. (2021). SegFormer: Simple and Efficient Design for Semantic Segmentation with Transformers. arXiv. https://doi.org/10.48550/arXiv.2105.15203
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2026 Margulan Nurtay, Gaukhar Alina, Ardak Tau

This work is licensed under a Creative Commons Attribution 4.0 International License.
The consolidation and conditions for the transfer of copyright (identification of authorship) is carried out in the License Agreement. In particular, the authors reserve the right to the authorship of their manuscript and transfer the first publication of this work to the journal under the terms of the Creative Commons CC BY license. At the same time, they have the right to conclude on their own additional agreements concerning the non-exclusive distribution of the work in the form in which it was published by this journal, but provided that the link to the first publication of the article in this journal is preserved.
A license agreement is a document in which the author warrants that he/she owns all copyright for the work (manuscript, article, etc.).
The authors, signing the License Agreement with TECHNOLOGY CENTER PC, have all rights to the further use of their work, provided that they link to our edition in which the work was published.
According to the terms of the License Agreement, the Publisher TECHNOLOGY CENTER PC does not take away your copyrights and receives permission from the authors to use and dissemination of the publication through the world's scientific resources (own electronic resources, scientometric databases, repositories, libraries, etc.).
In the absence of a signed License Agreement or in the absence of this agreement of identifiers allowing to identify the identity of the author, the editors have no right to work with the manuscript.
It is important to remember that there is another type of agreement between authors and publishers – when copyright is transferred from the authors to the publisher. In this case, the authors lose ownership of their work and may not use it in any way.